Downloading the PanSTARRS DR1 Catalogue via CasJobs
Before you begin running SQL queries on the PanSTARRS DR1 Catalogue via the command-line you first need to download the casjobs.jar java archive file and a config file.
Installation of CasJobs Command-Line Util
First move the jar files to a location on your machine’s $PATH. For example:
cd ~/Downloads
mv casjobs.jar /usr/local/bin/
Now create a directory somewhere where you’re going to be doing your work and move the config file into that directory:
mkdir ~/Desktop/panstarrs_dr1_casjob_batch_download
cd ~/Desktop/panstarrs_dr1_casjob_batch_download
mv ~/Downloads/CasJobs.config.x CasJobs.config
Note the removal of the .x extension on the config file.
Before we continue, you’ll need to find what your MAST WSID is. To find this ID, log into the MAST CasJob, navigate to ‘profile’ in the top toolbar and you should find your 9 digit WSID there:
Open the CasJobs.config configuration file in your favourite text-editor and amend the details to look like this:
wsid=<your_wsid>
password=<your_mast_password>
default_target=PanSTARRS_DR1
default_queue=1
default_days=1
verbose=true
debug=false
jobs_location=http://mastweb.stsci.edu/gcasjobs/services/jobs.asmx
To test your install run the command:
java -jar /usr/local/bin/casjobs.jar execute "select top 10 * from stackobjectthin"
If everything is setup correctly you should get 10 rows returned from the PanSTARRS StackObjectThin table. I added the following alias to my .bashrc file:
alias casjobs='java -jar /usr/local/bin/casjobs.jar'
so I can execute the same CasJob with the command:
casjobs execute "select top 10 * from stackobjectthin"
You can read more about PanSTARRS CasJobs here and a list of command you can execute with CasJobs here.
Select Which Data to Download
The PanSTARRS DR1 database has a few core tables containing the majority of the catalogued data and multiple helper tables and views that support those core tables. I’ve decided that I catalogued data I need are from the StackObjectThin and StackObjectAttributes tables.
Inspecting the PanSTARRS DR1 table schema from the CasJobs webpages we find that the StackObjectThin table contains \(\sim 3.5\times 10^9\) rows.
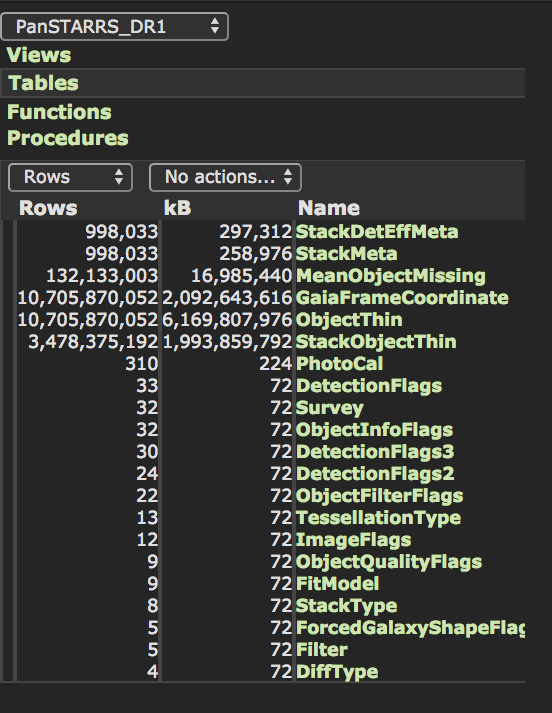
Select only rows with primary_detection = 1 leaves us with 2,877,925,753 rows to download. It’s also clear that StackObjectsAttributes is not a table but a view:
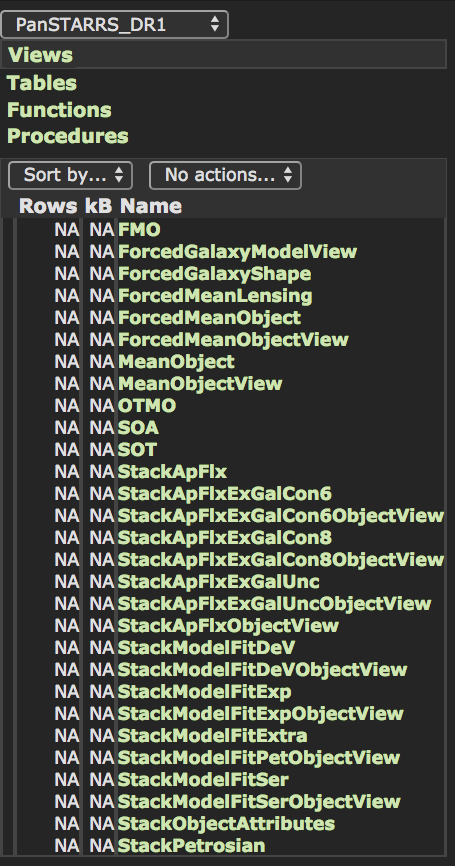
An Indexing Scheme for Systematic Downloads
Having targeted a few different combinations of columns for remote selection and indexing downloaded results, I 've settle on using just the objID column as I’m sure it’s used as a SQL Server index as MAST1. Also using a JOIN to combine the StackObjectThin and StackObjectAttributes table data resulted in very sluggish very times.
This script (see bottom of post) iterates over the StackObjectThin and StackObjectAttributes tables, ordered by the PanSTARRS objId and downloads packets of \(0.5 \times 10^6\) rows in FITS format. The filenames are prefixed with t for StackObjectThin and a for StackObjectAttributes and the 18 digit integer is the maximum objId in the file. For example:
t070002762982847811.fits
a070002762982847811.fits
Kicking off a Download
Before you run the download script, you need to make sure you have a spare 3TB of space on the machine you’re running the download on. As you can imaging \(\sim3 \times 10^9\) rows of data eats up a lot of bytes!
To run the script, download it into the same directory as your CasJobs.config file:
wget https://gist.githubusercontent.com/thespacedoctor/9c41f8417452719fba248ac3ddd4325c/raw/panstarrs_dr1_downloader.py
And run with:
python panstarrs_dr1_downloader.py
My initial speed tests are suggesting it should take 5–6 days to download all of the data.
-
using
projectionidandskycellidseemed like a smart idea but query times indicated it was going to take over 15 years for the download to complete! ↩
